Last Updated:
11th September 2020
About Vicente Martí Centelles
-

Vicente Martí Centelles
Vicente obtained a European PhD (Summa Cum Laude) from 2008 to 2012 in supramolecular chemistry with FPU fellowship from the Spanish Ministry of Science and Education at Univ. Jaume I working on the design and synthesis of macrocyclic compounds for molecular recognition. During his PhD he carried out two short research stays at Univ. Zaragoza (Spain, 2009) and Imperial College London (UK, 2010) obtaining a PhD prize from the Univ. Jaume I and preparing more than 10 publications, participate in more than 20 conferences, prepare one patent, participate in teaching at the university and pedagogic research projects (corresponding author of J. Chem. Ed. 2014 with more than 30 citations) and other soft skills acquired.
Then, he was awarded a 2-year postdoctoral VALi+d fellowship (2013–15) funded by the local government (Generalitat Valenciana, Spain) to develop a project involving University Jaume I, Universty of Oxford and a biotechnology company. Postdoctoral work allowed him interdisciplinary research, combining macrocyclic chemistry, interlocked hosts designed for molecular recognition and sensing of anions of biological and environmental importance and sensing of bacteria at the biotechnology company Biotica. This fellowship allowed writing one publication as the corresponding author (J. Org. Chem. 2016), a publication calcified as “Hot Paper” by the journal with an inside back cover (Chem. Eur. J. 2015) and a review on macrocyclization with more than 150 citations (Chem. Rev. 2015, IF=37.37).
The ensemble of these achievements allowed Dr. Martí to obtain the accreditation for assistant professor (2013) and the accreditation for associate professor (2015) from the Spanish National Quality Assessment Agency/Ministry of Education (ANECA). These are prerequisite qualifications for academic positions in Spain. Then he moved to work as Postdoctoral Research Assistant at the Univ. of Edinburgh in the design and synthesis of molecular machines and self-assembled host systems for molecular recognition and catalysis in the Lusby group. Dr. Martí discovered the catalytic properties of the Pd2L4 capsules, this key breakthrough shows an example of an artificial “Diels–Alderase” capsule that can overcome the product inhibition paradigm and combining efficient turnover alongside enzyme-like hallmarks (J. Am. Chem. Soc. 2018, IF=13.86, being Dr. Martí the main contributor). In addition to this paper, two more have been published (Chem. Sci. 2017, Chem. Eur. J. 2018) and others are under preparation. In January 2018 Dr. Martí obtained a prestigious Marie Curie IF fellowship (European Union funded) to continue his scientific career at the Nathan McClenaghan research group at the Institut des Sciences Moléculaires (CNRS/Université de Bordeaux) and Laurent Cognet research group focusing in supramolecular systems for fluorescence imaging. More details in the MSCA-IF project web page.
Dr. Martí publication years stared in 2011 and has published >30 papers, 3 of them as the corresponding author, with a total of 598 citations. His J. Am. Chem. Soc. 2018 paper has received 62 citations. The h-index is 12. See the publication list on his web or google scholar.
Awards and Fellowships obtained
- Marie Curie IF Fellowship from the European Union, 2018.
- Honourable mention in the XI Award Scientific-Technical "Ciutat d'Algemesí" for young researchers, 2015.
- Second prize in the XII Premios de innovación y creatividad Cátedra INCREA d'Innovació, Creativitat i Aprenentatge (Univ. Jaume I) for the pedagogical periodic table game ChemMend ( Chem. Ed. 2014), 2015.
- Postdoctoral VALi+d Fellowship from the Generalitat Valenciana, 2013.
- PhD Special Award basic sciences (Premio extraordinario de doctorado en ciencias básicas) Univ. Jaume I, 2013.
- Chemistry Bachelor Honourable Mention in the Chemistry the National Awards for University Studies (Premios Nacionales Terminación Estudios Universitarios) Spain Education Ministry, 2006/7.
- Predoctoral FPU Fellowship from the Spanish Ministry of Science and Education, 2008.
- Chemistry Bachelor Award (Premio extraordinario final de carrera Lic. Química) Universitat Jaume I, 2006/7.
- Chemistry Bachelor Award (Premio al rendimiento académico Generalitat Valenciana), 2006/7.
- Chemistry Bachelor Award Premio Ernest Breva Mallach al esfuerzo y excelencia en los estudios de licenciatura en Química (School of Technology and Experimental Sciences), Universitat Jaume I, 2006/7.
- Chemistry Bachelor Award Premio Colegio de Químicos CV al mejor expediente académico, 2006/7.
- Intitiation to research Fellowship from the Spanish Ministry of Science and Education, 2006.
- Chemistry Bachelor Award Premio al Rendimiento Académico Primer Ciclo Lic. Química, Univ. Jaume I, 2002/5.
- Silver Medal in the 35th International Chemistry Olympiad, Groningen, The Netherlands, 2002. The highest distinction obtained by Spain in the International Chemistry Olympiad.
- Chemistry Olympiad Silver Medal at 7ª Olimpiada Iberoamericana de Química, Mar del Plata, Argentina, 2002.
- Chemistry Olympiad Gold Medal in the 13ª Olimpiada Nacional de Química, Spain, Oviedo, 2002.
- Physics Olympiad Honourable mention in the 13ª Olimpiada Española de Física, Spain, Burgos, 2002.
- Chemistry Olympiad Gold Medal in the Olimpiada de Química año 2002, Universitat Jaume I, 2002.
- Physics Olympiad Gold Medal in the XIII Olimpiada de Física, Universitat Jaume I, 2002.
Research
Vicente Martí Centelles research interests are focuses in the applications of supramolecular systems in different fields. Recent research is focused in the design of macrocycles, molecular cages, interlocked molecules, etc. with tailored properties ranging from molecular sensing, catalysis and microscopy.
MSCA-IF: Supramolecular Chemistry for super-resolution microscopy


Vicente Martí Centelles current research interests focuses on supramolecular systems for advancing the field of fluorescence imaging. This research is carried out through his Marie Curie IF fellowship in the Nathan McClenaghan research group at the Institut des Sciences Moléculaires (CNRS/Université de Bordeaux) and Laurent Cognet research group at the Laboratoire Photonique Numérique et Nanosciences - Institut d'Optique Graduate School (CNRS/Université de Bordeaux). Super-resolution microscopy allows obtaining high resolution images overcoming the light diffraction limit. Thanks to the Marie Curie Individual Fellowship that I obtained, I am developing fluorescent supramolecular systems able to work efficiently in this microscopy techniques. CORDIS EU web page of the MSCA-IF project https://cordis.europa.eu/project/rcn/215464. More details in the MSCA-IF project web page.
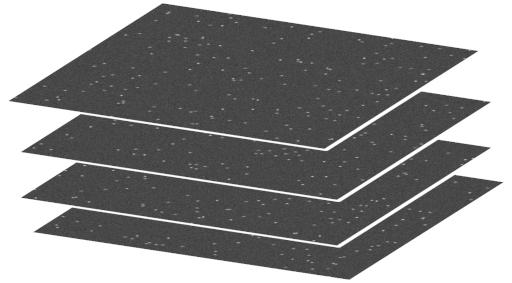

This project, indeed allows to integrate all state of the art knowledge acquired during my career to develop functional molecules with tailored properties in microscopy. It is fascinating to be able to detect single molecules using microscopy. This allows studying the individual behavior of molecules that cannot be distinguished using bulk measurements, where it is only possible to measure average values. Single-molecules detection can be performed in a confocal microscope using appropriate fluorescent molecules, excitation laser and a camera. The following image shows differnt frames containing single molecules simulted with ThunderSTORM in ImageJ, and ilustrates the process of detecting single molecules over time to reconstruct a high resolution image by processing all the individual frames. In an interview with the "Sociedad para el Avance Científico" to talk about my Marie Curie Fellowship, you can read the full interview and watch the Youtube video to get further details about the process of getting a Marie Curie Fellowship, the benefits from it and an overview of the project objectives.

-
Entrevista al Dr. Vicente, receptor de una beca Marie Curie
SACSIS | Dic 2, 2019 Leer entrevista completa
Artificial enzymes
Vicente Martí Centelles was previously a postdoctoral researcher in the Lusby group
at the University of Edinburgh.
His research focused in the design and synthesis of molecular machines and self-assembled host systems for molecular recognition and catalysis.
Dr. Martí discovered in his research the catalytic properties of the Pd2L4 capsules, and he developed experimental protocols and kinetic
models to obtain quantitative thermodynamic data by means of data science programing.
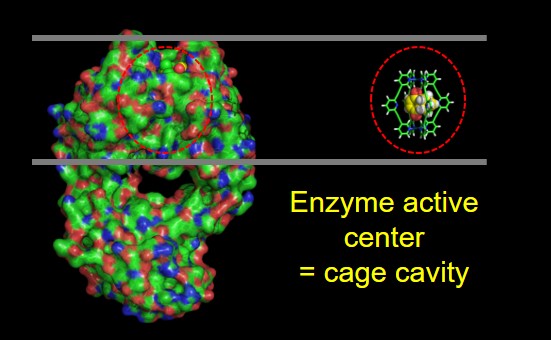
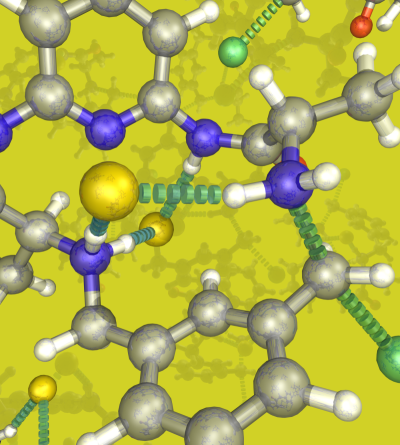
The catalytic hosts have properties similar to the biological systems, in particular, the proteins. The unfolded peptides do not have any activity, and the folded structure has a very efficient activity for a particular reaction. For the cage compounds, a similar effect is observed. The disassembled components do not have any catalytic activity, but the assembled structure has enhanced molecular recognition and catalytic properties. So the catalysis is an emergent property of these sophisticated molecular architectures. Focusing on the design of the cages, why are enzyme active sites buried? If we analyze the size of the active center of an enzyme in comparison to all the structure, this is relatively small with similar size of our cage design. The most important supramolecular interactions present in an enzyme active center are multiple hydrophobic contacts providing selectivity and multiple non-covalent interactions being the main contribution to activity and also providing selectivity.
This research resulted in a key publication J. Am. Chem. Soc. 2018, 140, 2862-2868. IF=13.86, being Dr. Martí the main contributor) reporting an artificial "Diels-Alderase" capsule overcoming the product inhibition paradigm and combining efficient turnover alongside enzyme-like hallmarks. Along the different research topics, Vicente has been involved in the organization of conferences, and being a member of the organizing committee the 3rd edition of the EaStCHEM Conference for Early-Career Researchers ECECR 2018 that was at King's Building Campus, University of Edinburgh.
-
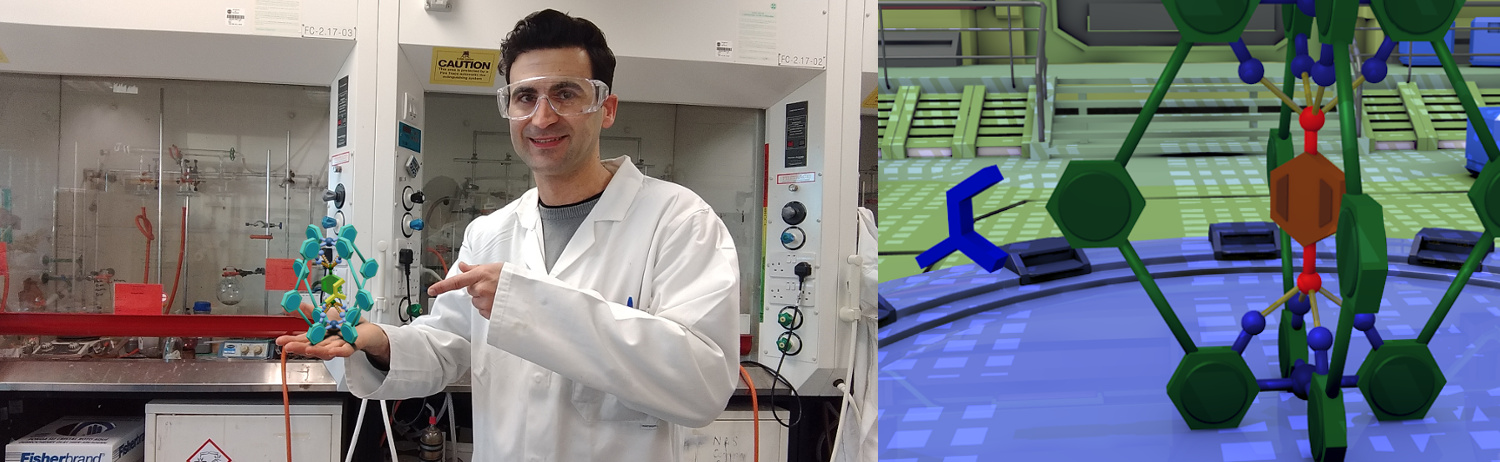

High Activity and Efficient Turnover by a Simple, Self-Assembled Artificial Diels-Alderase
J. Am. Chem. Soc. 2018, 140, 2862-2868. J. Am. Chem. Soc. 2018, 140, 2862-2868. -
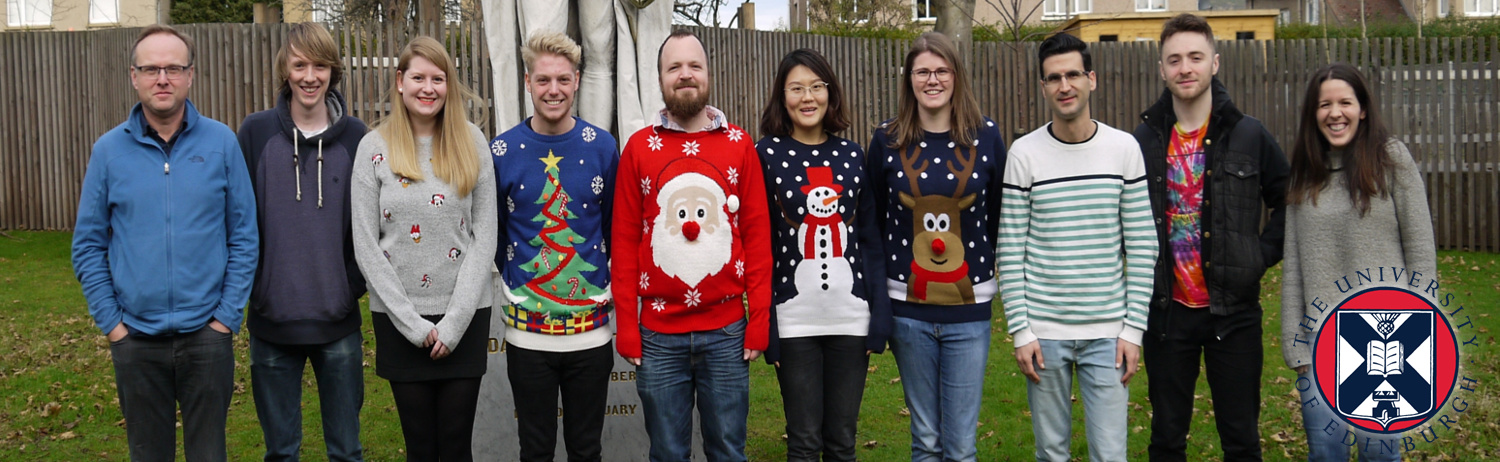

The Lusby Group - University of Edinburgh
The Lusby Group - University of Edinburgh - 2017 www.lusby.chem.ed.ac.uk -
An "Artificial Diels-Alderase" in action
View video in Youtube
Interlocked systems for molecular recognition
Postdoctoral work through the VALi+d fellowship allowed Vicente to diversify his research activities, focusing on macrocyclic chemistry at Univ. Jaume I, synthesis of interlocked hosts designed for molecular recognition and sensing of anions of biological and environmental importance at Univ. of Oxford (Chem. Eur. J. 2015, “Hot Paper”, inside back cover) and sensing of bacteria at Biotica. All the knowledge acquired allowed publication of a review on macrocyclization (Chem. Rev. 2015), and additional participation in two general scientific magazine publications.
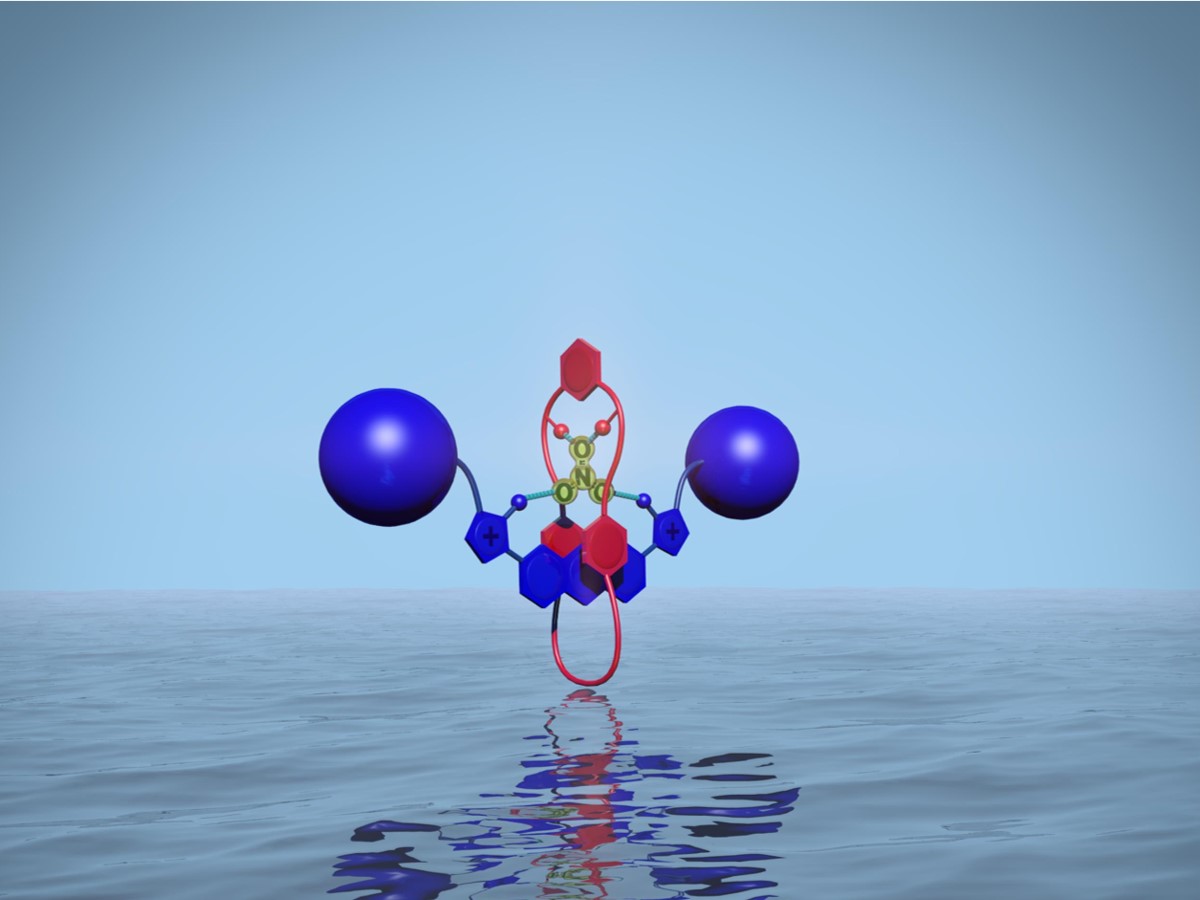

The work developed at the University of Oxford focused on the design and preparation of receptors of species of biological, environmental, and medical interest. In this sense, the essential role of anions in these fields must be emphasized. The important role of anions has motivated chemists to develop synthetic anion receptors capable of selective recognition, this research area being an important part of the field of Supramolecular Chemistry.
Nitrate anion plays an important role in environmental and medical contexts: excessive use of fertilizers in agriculture has caused eutrophication and alteration of the natural aquatic ecosystem, while exposure to elevated nitrate levels is a cause of diseases such as methemoglobinemia (blue baby syndrome) in infants. The preparation of selective nitrate receptors is a poorly developed area, with few examples described in the literature, and which mostly operate only in organic solvents. The main challenges to overcome in the design of selective nitrate receptors are the low affinity of the anion to form hydrogen bonds and the high hydration energy of the anion in mixtures of aqueous solvents, which contribute to an association weak in general with the host molecule.
These results were hihghlighted in the news through several interviews. Vicente Martí-Centelles tells to the journal Pan European Networks: Science & Technology about the development of a pioneering method in the evaluation of nitrate pollution Pan European Networks: Science & Technology June 2016, From the UJI to Oxford against water pollution Mediterraneo 22/10/2015, a researcher creates a method that detects nitrates selectively Levante de Castelló 22/10/2015, a researcher designs a pioneering method in the nitrate pollution assessment Universitat Jaume I Science Videos, interview on Radio Voramar esRadio 92.5,in "Es la Mañana" to explain nitrate detection research Radio Voramar esRadio 92.5 29/10/2015, dissemination of research on nitrate detection in Finestra a la Ciència de Vox UJI Radio 30/10/2015, Blog Ciencia TV Universitat Jaume I 2015, and Universitat Jaume I web page 2015.
Whit this work I was able to obtain a honourable mention in the XI Award Scientific-Technical "Ciutat d'Algemesí" for young researchers in 2015. This was hihglhted in the news Algemesi rewards the research work on environmental pollution carried out by Vicente Martí.
Macrocyclic chemistry
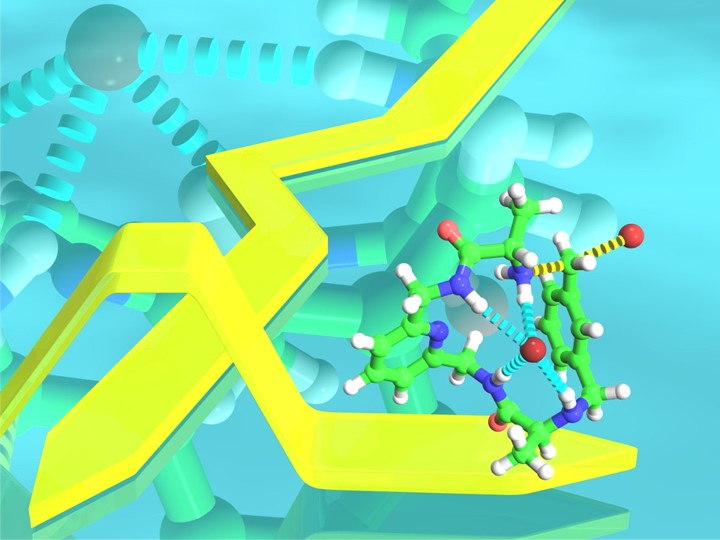
Vicente PhD thesis work focused on the design and preparation of pseudopeptidic systems with applications in molecular recognition and self-assembly. Vicente developed this work at Univ. Jaume I, and performing two short research stays in leading research groups at Imperial College London and Univ. Zaragoza in order to gain experience in specific research topics. Vicente developed anion template synthetic strategies to achieve an efficient synthesis of macrocyclic structures through SN2 reactions in the macrocyclization reaction step (Chem. Eur. J. 2012, J. Org. Chem. 2014, Sci. World J. 2012). Some of these macrocyclic compounds based on acridine had excellent anion recognition and sensing properties, and they were able to selectively recognise dihydrogen phosphate and the tryptophan amino acid by means of fluorescence experiments (Chem. Eur. J. 2014, J. Org. Chem. 2012). In his PhD work he also prepared and characterized novel polymeric metal complexes with appropriately designed pseudopeptidic ligands in my research stay at Imperial College London (CrystEngComm 2011). In these scientific publications he was the main contributor and 1st author. Additionally, he did further studies in the topic of anion-templated macrocyclization reactions in my postdoctoral project at Univ. Jaume I. In this case, Vicente prepared the macrocycles by means of amide bond formation reactions in the macrocyclization step (J. Org. Chem. 2016). During the PhD, Vicente also participated in a project where a family of organogellating compounds were able to form gels by the self-assembly of the monomers and the gels were stable to high temperatures, these studies resulted in the preparation of one publication (Tetrahedron 2013) and one patent.
Vicente Martí Centelles research in supramolecular chemistry exceptional combination of skills: organic chemistry, supramolecular chemistry, and physical organic chemistry.
-

Vicente Martí Centelles - Graphical Abstract Publications
Vicente Martí Centelles Publications
Dissemination of research results and outreach activities
Vicente Martí Centelles has participated in different outreach activities for disseminating his research work targeting a broad audience. These activites include, the european researcher night 2019 in Bordeaux, the "Fete de la Science 2019" at Institut de Sciences Molecularies in Bordeaux and preparing with journalists press releases and journal/radio/video interviews.
-
Entrevista al Dr. Vicente, receptor de una beca Marie Curie
SACSIS | Dic 2, 2019 Leer entrevista completa -

European researcher night and Fete de la Science
Vicente Martí Centelles, Bordeaux 2019 View larger image -

Inteview in Pan European Networks: Science&Technology
Pan European Networks: Science&Technology, June 2016 Read complete interview -
An "Artificial Diels-Alderase" in action
View video in Youtube -
Dissemination of the nitrate detection
Univ. Jaume I Science Videos Youtube video -

A method that selectively detects nitrates
Levante de Castelló - 22/10/2015 Read complete article -

From the UJI to Oxford against water pollution
Mediterraneo - 22/10/2015 Read complete article -

Radio interview about nitrate detection
Radio Voramar esRadio 92.5 - 29/10/2015 Complete interview -

Radio interview about nitrate detection
Finestra a la Ciència de Vox UJI Radio, 30/10/15 Complete interview -

Nitrate detection - Blog Ciencia TV Universitat Jaume I
Blog Ciencia TV Universitat Jaume I - 2015 Complete interview -

Nitrate detection - Universitat Jaume I web page
Universitat Jaume I web page - 2015 Complete interview -

Organogelating compunds patent outreach activities
Universitat Jaume I web page - 2011 Complete interview
International Chemistry Olympiad
Vicente Martí Centelles has participated in the Chemistry Olympiad over the last years. Dr. Martí's interest in Chemistry began early and in 2002 he obtained a gold medal in the Spanish National Olympiad and a silver medal in both the International Chemistry Olympiad (the highest distinction obtained by Spain in this prestigious competition) and Ibero-American Chemistry Olympiad. The participation in the Chemistry Olympiad includes:
- Member of the Scientific Committee Univ. Jaume I Chemistry Olympiad (2009-13, and 2015).
- Member of the scientific committee of the Spanish National Chemistry Olympiad. 2013 (Alicante), 2012 (Madrid), 2011 (Valencia), 2010 (Seville), 2009 (Avila).
- Head Mentor (head of the Spanish delegation, 2015 Azerbaijan), Mentor (2011 Turkey, 2012 USA, and 2013 Russia) and scientific observer (2009 UK, 2010 Japan), collaborator in training (2008) of the Spanish delegation at the International Chemistry Olympiad (IChO).
The training carried in the Chemistry Olympiad as mentor, cientific observer and collaborator has been succesful, and the students have obtained bronze medals and honourable mentions every year:
- 2008 BUDAPEST (HUNGRÍA): Eduardo Ansaldo Giné, Medalla de Bronce.
- 2009 CAMBRIDGE/OXFORD (INGLATERRA): David Rodriguez San Miguel, Medalla de Bronce; Luca Schneller-Pavelescu, Mención de Honor.
- 2010 TOKIO (JAPON): Pablo Giomi, Medalla de Bronce Jesús Álvaro Gómez Iregui, Medalla de Bronce; Andreu Tortajada Navarro, Medalla de Bronce.
- 2011 ANKARA (TURQUIA): Moisés Maestro López, Medalla de Bronce; Albert Soto Company, Mención de Honor.
- 2012 WASHINGTON (E.U.A.): Sergio Tomás Martínez, Medalla de Bronce.
- 2013 MOSCÚ (RUSIA): Damiá Torres Latorre, Medalla de Bronce; David Prieto Rodríguez, Medalla de Bronce; Sergio Cuesta Galisteo, Mención de Honor.
- 2015 BAKÚ (AZERBAYÁN): Raimon Terricabres Polo, Medalla de Bronce; Jorge Melendo Arrufat, Mención de Honor.
-

IChO Spanish Team
View larger image -

IChO 2015, Baku, Azerbaijan, Spanish Team
View full size image -

IChO 2013, Moscu, Russia, Spanish Team
View full size image -
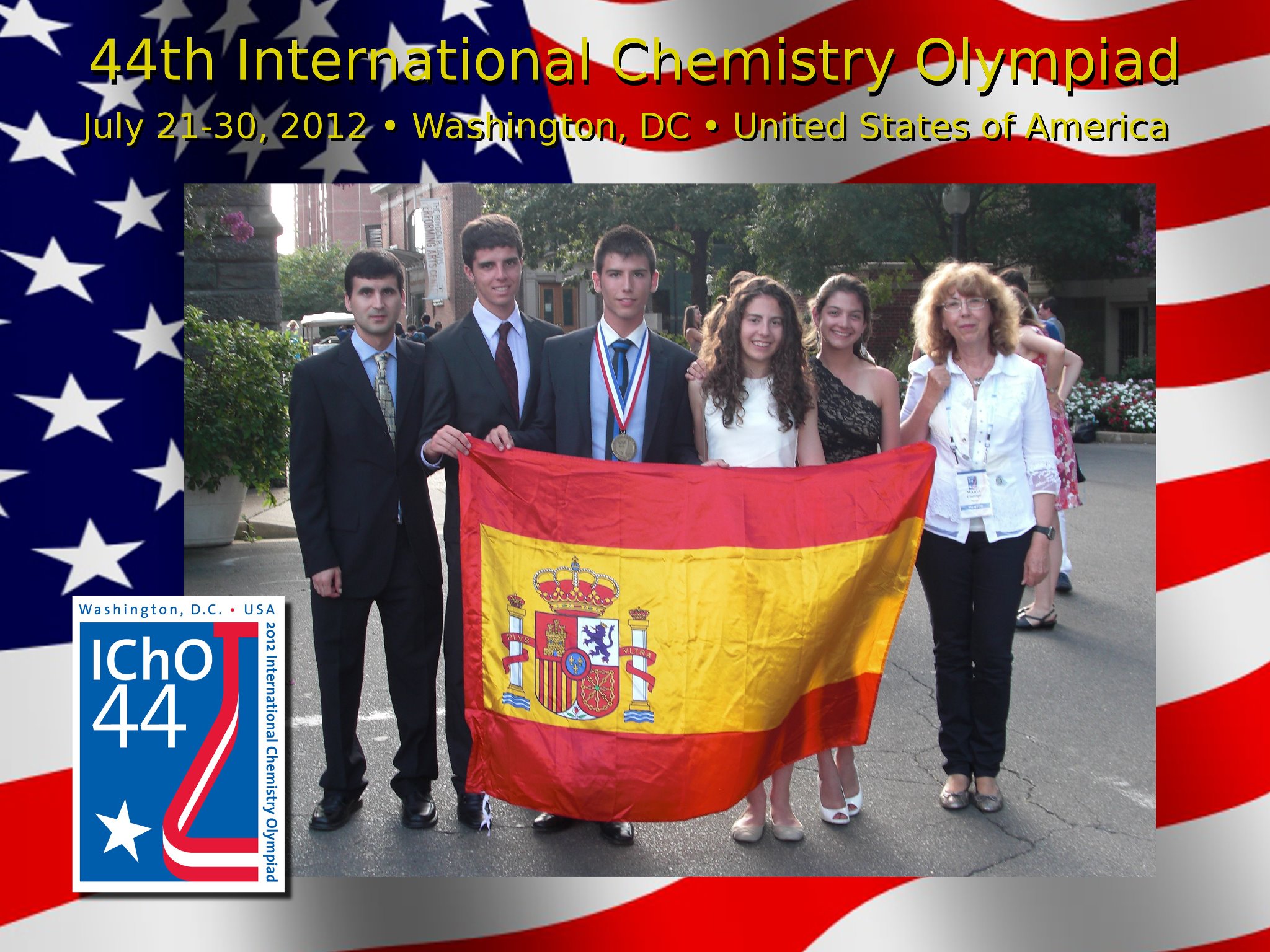

IChO 2012, Washington, USA, Spanish Team
View full size image -

IChO 2011, Ankara, Turkey, Spanish Team
View full size image -

IChO 2010, Tokyo Japan, Spanish Team
View full size image -

IChO 2009, Oxford & Cambridge, UK, Spanish Team
View full size image -
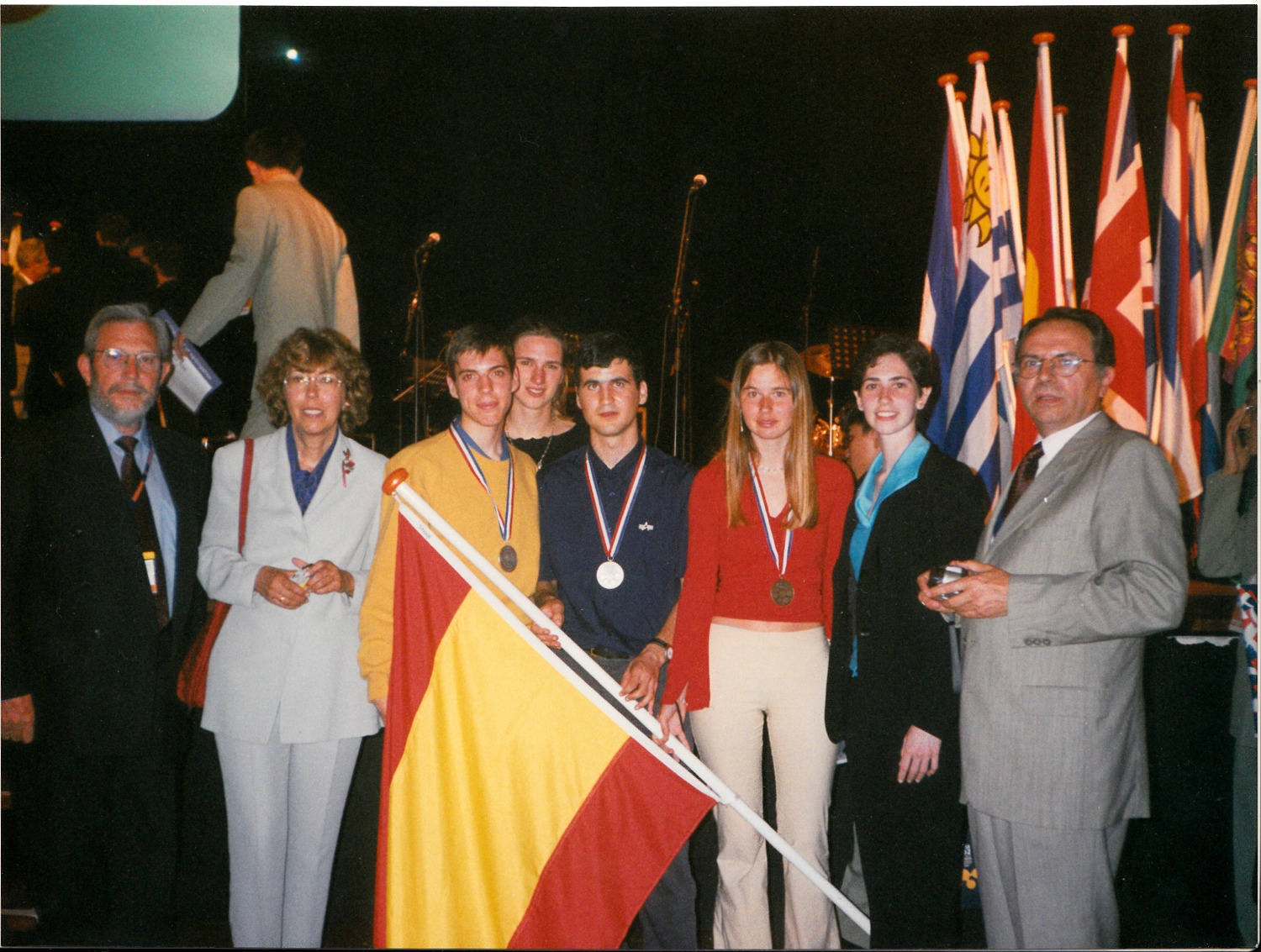

IChO 2002, Groningen, The Netherlands, Spanish Team
View full size image -

IChO International Chemistry Olympiad Flag
View full size image
Vicente whon a Silver Medal at the International Chemistry Olympiad in 1002, and this was highlighted in the news, a complete press dossier is available with all the interviews.
-

Press Dossier - Silver Medal at the International Chemistry Olympiad 2002
Vicente Martí Centelles - IChO 2012 Complete press dossier -

Speech at the awards ceremony of the Science Olympiad 2010/2011 at Universitat Jaume I
Vicente Martí Centelles - Universitat Jaume I Read complete speech
ChemMend: Learning the periodic table with a card game

Knowing the Periodic Table is essential in the field of chemistry, but its memorisation is often tedious and complex for students. Vicente Martí Centelles and Jenifer Rubio Magnieto, alumni of Chemistry at the Universitat Jaume I, have developed a game for secondary school students and first years of university, and anyone interested in chemistry, to learn the group and period each chemical element belongs to in the Periodic Table in an easy, simple and funny way. Two years of work and experimentation in the classroom have resulted into ChemMend, a game that has been reported on the International Journal of Chemical Education.
-

The Jaume I supports, through UJI Emprèn, the launch of the game to learn the Periodic Table, ChemMend
Universitat Jaume I - 2015 Read the news
ChemMend is the first existing game that allows exploring and gaining knowledge about the groups and periods of elements since; so far, existing games on the Periodic Table are focused on linking the different elements with their symbol. The gameplay is similar to the popular UNO, but this time is to follow the reference card with cards that correspond to the same group of elements or the same period, or to use one of the cards that act as a wildcard and which introduce different instructions as jumping a turn, draw cards from the deck, change direction, make a drawing, etc. All the info can be found in the ChemMend Full Press release document. Also you can found further infomration on www.facebook.com/ChemMend the orignal publication J. Chem. Ed. 2014, 91, 868-871. and the electronic online implementation. www.chemmend.uji.es.
Software
-
Augmented reality for chemistry
Vicente Martí Centelles View video in Youtube
Development of Chemistry Software
I've developed software during my different research process. At the moment I am working on my Vicente GitHub site to share the soruce code, and this sofware will be available to download in due course. I have learned different programming languages that allowed to develop these software. Such languages include C++, Python, Cran-R and Mathematica.
I am also develping a tool to improve the presentation of complex strucutures in poster presentations or meetings. This tool is java script based, and it does not requires the installation of any app and allows displaying a molecule over the AR.Chem recognition motif. The main advantage of this methodology is to allow the user to acces directy to the 3D molecule just using an smartphone to read the QR code, then a web page will be oppened asking to acces the camera and the augmented reality molecule will be displayed over the QR code, simple and efficient. The source code can be accesed from my GitHub page on the following link: Augmented reality for chemistry
You can simply test the this code by acccesing the following link, and then point the camera of you smartphone to the image below: Run the augmented reality for chemistry in your smartphone.
HTML code to implement AR.Chem

<!doctype HTML>
<html>
<script src="aframe.min.js"></script>
<script src="aframe-ar.js"></script>
<script src="aframe-vrml-component.min.js"></script>
<body style='margin : 0px; overflow: hidden;'>
<a-scene embedded
vr-mode-ui="enabled: false"
arjs='debugUIEnabled: false;
sourceType: webcam;
detectionMode: mono_and_matrix;
matrixCodeType: 3x3
trackingMethod: best;'
renderer="antialias: false;
logarithmicDepthBuffer: true;
colorManagement: true;
sortObjects: true;"
>
<a-assets>
<a-asset-item id="3d-wrl" src="rotax_v01.wrl"></a-asset-item>
</a-assets>
<a-marker type='pattern' url='ar-chem.patt'>
<a-entity
vrml="#3d-wrl"
scale ="0.15 0.15 0.15"
position="0 0.5 0"
rotation="-90 0 0"
>
</a-entity>
</a-marker>
<a-entity camera></a-entity>
</a-scene>
</body>
</html>
If you are interested in using any of these contact me by e-mail and I can provide it or we can collaborate to implement it you your particular needs or to solve any particular challenge that you have in your research. For example, I have developed a wide variety of Advanced Kinetic Modelling code with R that allows simulating very complex reactions and perform multivariate fitting.
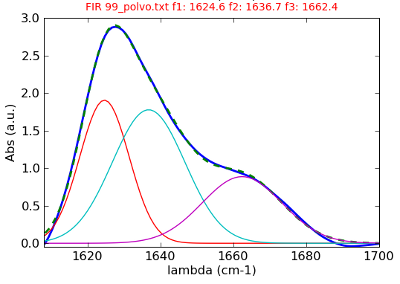
These sofware has been developed in the Python programming language (using the libraries: SciPy, Matplotlib and PIL Python Image Library) and R. Some of these programs had an user graphical interface (GUI) develped by using the GTK interface wit the aid of glade.
- SEM batch image processing
- Batch image processing to put all the images labeled into the same image and with the correct label
- IR-ATR deconvolution and fitting
- Advanced Kinetic Fitting with R
- UV and CD script to batch process files
- Advanced batch processing of xyz files to create Gaussian input files
- Script to automatically represent a PES from a Gaussian calculation
High Quality Renders for Chemical Structures
We can create high quality renders for our chemical models with Blender. To do this render first we need a pdb file of our molecule. I usually use gaussian to perform my calculantions and I use molden to view the output files. Then I export the molecule in Tinker xyz format. Then I convert the tinker xyz file to pdb wiht xyz2pdb ( the xyz2pdb-src-linux_bin-windows_bin.zip contains the source code (written in C) and binaries compided for linux and windows). To use xyz2pdb open a terminal in linux or the cmd in Windows and execute xyz2pdb -help to obtain a list of help:
$ ./xyz2pdb -help
Usage: ./xyz2pdb [<flags> with arguments, as follows]
-o <PDB file name> default is <stdout>
-xyzfile <XYZ file name> default is <stdin>
-debug <level> 0 to 9 [0]
-backbone output backbone only
-noh don't output hydrogen atoms
-noh2o don't output water molecules
-nourea don't output urea molecules
-usebond derive bond info from input file only
-help print this information
Therefore to convert our xyz file we must use:
$ ./xyz2pdb -xyzfile myfile.xyz -o myfile.pdb
Then we will import this pdb file into Blender with pdb2blender.py, this python script is already in a blender file, pdb2blend12.blend. To use the script open the file pdb2blend12.blend in blender, then in the left window right click and execute script, setup the parameters to import and click on Import button to select the pdb file.
Now we can play with blender and modify the properties of the materials, add some reflections... (you can try with this files: ts-tinker.xyz, ts.pdb).
Vicente Martí Centelles - 2019